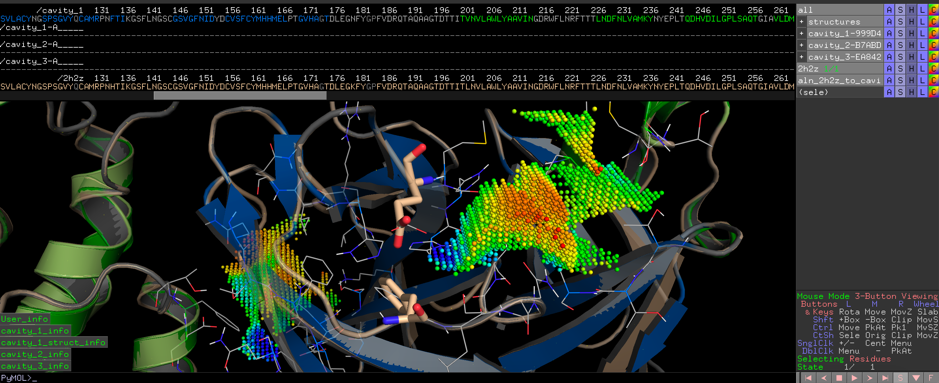
Coronavirus COVID-19 (formerly known as Wuhan coronavirus and 2019-nCoV) - what we can find out on a structural bioinformatics level - Innophore
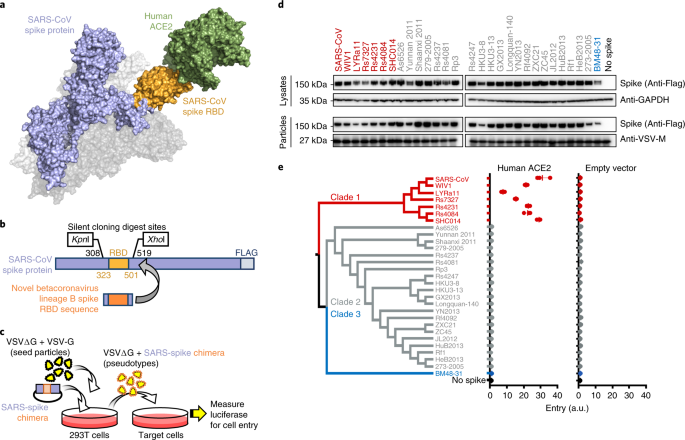
Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses | Nature Microbiology
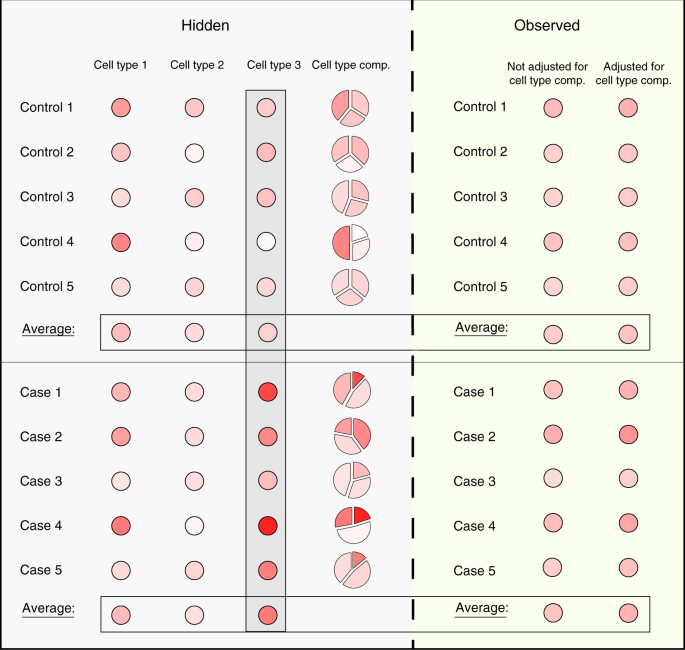
Cell-type-specific resolution epigenetics without the need for cell sorting or single-cell biology | Nature Communications


![Underrepresentation of women in the senior levels of Brazilian science [PeerJ] Underrepresentation of women in the senior levels of Brazilian science [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2017/4000/1/fig-1-2x.jpg)